February 19, 2019
By Peter Newman
Alexander Graham Bell famously said,
“When one door closes another door opens, but we so often look so long and so regretfully upon the closed door, that we do not see the ones which open for us.”
Atrazine resistant wild radish looks a lot like a door closing, but in many cases, it’s also a door opening for Bromoxynil.
Some new research by AHRI PhD student, Huan Lu, has shed some light on Atrazine resistant wild radish and the results have some very practical applications for growers and agronomists.
The most common target site mutation that causes Atrazine resistance in wild radish is the 264 mutation, the same mutation that gave us TT Canola. The only way to kill radish with this mutation is to sit the Atrazine drum on top of them! They have very high resistance to Atrazine and as it turns out they are super-sensitive to Bromoxynil, requiring only about a third of the Brom rate that it takes to kill susceptible wild radish. Wild radish with the 264 mutation is also resistant to metribuzin.
Huan Lu also discovered a new target site mutation, the 274 mutation that causes modest levels of resistance to Atrazine, Metribuzin and Diuron but not Bromoxynil.
If you are gazing regretfully at Atrazine resistant wild radish perhaps spare a thought for the super sensitivity to Bromoxynil that may have resulted.

Wild Radish. Photo by: Danica Goggin.
Atrazine resistant radish is easy to kill with Bromoxynil
This is big news, and you may have even seen it with your own eyes in the field. Huan Lu found that if you have highly resistant radish with the very common 264 mutation it’s super sensitive to Bromoxynil. This phenomenon is also sometimes called negative cross-resistance. Wild radish with the 264 mutation has an LD50 for Bromoxynil of only 15g/ha compared to 49g/ha for the susceptible (Table 1). In other words, it’s about three times as easy to kill with Bromoxynil. Products such as Jaguar® and Velocity® that contain Bromoxynil get a free kick with triazine resistant radish.
Wild radish with the less common 274 mutation was not resistant to Bromoxynil.
How does it work?
Atrazine binds to the D1 protein by Hydrogen bonds. When the 264 mutation is present, Atrazine can’t form these H-bonds and high-level resistance results. Bromoxynil is the complete opposite. When the 264 mutation is present, bromoxynil forms an extra H-bond to the D1 protein resulting in super-sensitivity to Bromoxynil. The D1 protein is an integral part of the electron transfer in photosynthesis.

Table 1: LD50 (lethal dose in gai/ha to kill 50% of the population) of a susceptible wild radish population compared to wild radish with either the 264-Gly or 274-Val mutation. Two to three leaf wild radish.
The 274 mutation
This research discovered the first evidence of the 274 mutation in wild radish. This mutation causes modest levels of resistance to Atrazine, Metribuzin and Diuron as can be seen in the table above and the dose-response curves below.
In Australia, the triazines (eg. Atrazine), the triazinones (eg. metribuzin), the Ureas (e.g. Diuron) and the Nitriles (e.g. Bromoxynil) are all group C herbicides. They all target photosynthesis, more specifically the electron transfer in PSII, but at slightly different binding sites, so we can’t treat them all the same. There are seven known target site mutations that cause resistance to this group of herbicide.
Dose-response
The dose-response curves below illustrate the difference in resistance patterns between the different mutations. It’s interesting to see how highly resistant the 264 population is to Atrazine in chart (a), yet the opposite is true for Bromoxynil in chart (d).
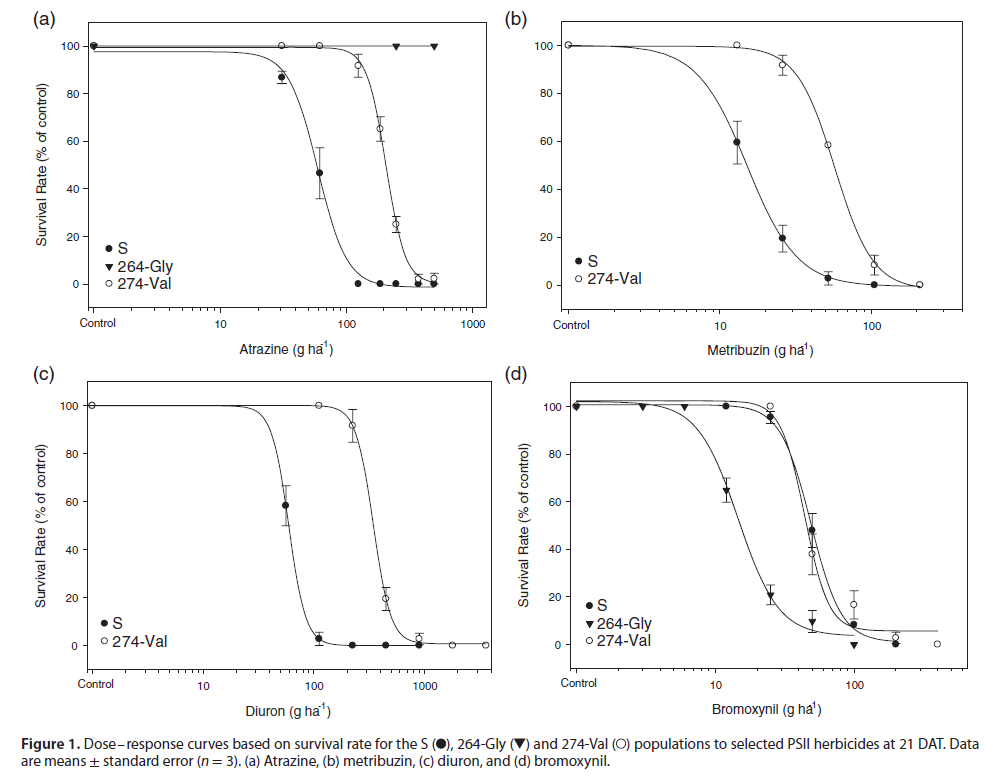
Metribuzin and the 264 mutation
You’ll notice in the dose-response curves above that Huan Lu didn’t do a dose-response for metribuzin for the 264 population. Other research has confirmed that the common 264 mutation also causes metribuzin resistance, but not the very high-level resistance that we see with atrazine with this mutation. This may explain why some farmers and agronomists have seen that they still get some useful control from metribuzin on wild radish in lupins where they know they have atrazine resistant wild radish.
The Northern Ag region of WA
The northern wheatbelt of Western Australia represents Australia’s highest level of adoption of glyphosate-tolerant canola with an approximate 70% market share. Why? Triazine-resistant wild radish. This region is famous for the very long lupin-wheat rotation that some farmers have now operated for 40 years or so. This rotation is heavily reliant on simazine, and it’s likely that this long period of simazine use resulted in the widespread evolution of triazine resistant wild radish in the region. The only way to grow the profitable break crop, Canola, in this region for many grain growers is to use the glyphosate-tolerant technology to enable successful wild radish control.
Breeding of TT canola
An interesting side note to this research is the story of how triazine tolerant (TT) canola was bred. A breeding program in Canada crossed a brassica weed with the 264 mutation with canola to produce TT canola. We know that this causes a 30% fitness penalty in canola and it is likely that the same will occur in wild radish, but given that wild radish is already exceptionally fit we may not notice it in the field.
Summary
At times it can be hard to see where all of this molecular biology research is headed, however, this research can also give us great insight and help us understand what we are seeing in the field. This deep understanding of herbicide resistance mechanisms can have very practical applications and go a long way to helping us make good weed control decisions.
Paper
Posted in: Herbicide resistance mechanisms


